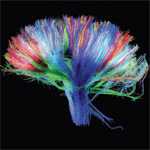
 Successful replication of a mature human brain – one that we would accept as a replacement for our own, or for the brain of someone we love – must preserve almost all the connections within it. Connections embody the psychological properties that make human individuals who they are: memories, learned abilities, habits, associations, talents, and emotional responses.
Successful replication of a mature human brain – one that we would accept as a replacement for our own, or for the brain of someone we love – must preserve almost all the connections within it. Connections embody the psychological properties that make human individuals who they are: memories, learned abilities, habits, associations, talents, and emotional responses.
In an organ such as the liver, it doesn’t matter that two particular cells are adjacent, because the liver is not a communications network. In the brain, the physical arrangement of individual cells matters very much. All our psychological attributes – the differences between the minds of an Einstein and a Hitler – are instantiated in that physical relationship.
Information Estimate for the Human Brain
A 2009 study by Azevedo et al found that the adult human brain contains, on average, 86 billion neurons and 85 billion neuroglial cells, or glia. Although most neurological research focusses on neurons, glia are equally vital to proper brain function. They provide physical support to neurons, supply them with nutrients and oxygen, destroy pathogens, remove toxins and dead material. They are also directly involved in the brain’s circuitry, supplying the myelin sheath responsible for electrical insulation in the brain, and modulating neurotransmission. They only lack action potential – the ability to transmit signals in the brain. Glia are so important that it is prudent to assume that the degree of completeness and accuracy required for glial replication is about equal to the standard required for neural replication. The precise location of blood vessels may also be more important in the brain than in other organs. Until we know better, we should apply the same standard to them too.
That standard is a high one. When it comes to replicating a brain, unlike other organs, we cannot afford to generalize, ignoring individual cells. A great deal of information about each neuron must be recorded and faithfully transferred to the replica. How much information? To begin with, the replication process should aim to preserve all synaptic connections. Wikipedia cites an article by Drachman estimating an average of 7000 connections per neocortical neuron in adults. (Young children have many more. More connections are pruned away than created during maturation of the brain!) The identities of the two neurons involved in each synaptic connection are certainly important. A unique number identifying a single neuron in a population of 86 billion can be expressed in 37 bits of information. To identify the two neurons would take 37 + 37 = 74 bits per connection, or 518,000 bits (65 kilobytes) per neuron. Multiplying by 86 billion neurons gives a total of 5.59 petabytes (PB) of information. That’s just for the basic connectivity map: a record of which neurons are connected to which. More information would be required.
We would also need to know the type of synaptic connection (whether electrical or chemical, and if chemical, the specific neurotransmitter to which the synaptic receptor responds). As that is largely a function of the transmitting neuron, it might not need to be expressed as a property of the connection; but if it were, then estimating 100+ neurotransmitters, that would take another 9 bits per connection. The 3D spatial location of the synapse is also important; it could be expressed to 1 nm precision (probably overkill) using 93 bits. Therefore we could express the type and location of each neural connection, and the identities of the transmitting and receiving neurons, in 74 + 9 + 93 bits = 176 per connection. That multiplies out to over 13 PB for the whole brain.
The careful reader may have noticed that this exceeds my earlier estimate of 1 petabyte of information needed to replicate the entire human organism. That was based on assumptions that now appear too rough. Lumping all human cells together in terms of information content, I allowed 16 bits for cell type and 128 bits for spatial position and shape, for a total of 144 bits per cell. Given current scientific progress in tissue engineering, that now appears to overestimate the information required for most somatic cells, but vastly underestimate what is needed for neural cells, especially the neurons of the brain.
To produce a ‘fair copy’ of a human being, we don’t need details about individual cells belonging to most of the body’s tissues. We need a lot of details about the individual cells of the brain. That suggests that of all the information which would have to be sent over the internet in order to teleport me to Omaha, the volume of data devoted to my brain would exceed the data representing the rest of my body by many times. Although my brain weighs just 2% of my body, the ‘informational weight’ of my brain – dominated by the connectivity map – might well come in at 95% or higher!
Let’s work with those numbers. Remember, the 13 PB estimate is only for neural connectivity. Other properties of the neurons are important, as are attributes of the glia and blood vessels of the brain. But the volume of information required to capture those attributes is on the level of whole cells rather than connections; as such it is insignificant compared to the connectivity information. One more petabyte should be enough. Accordingly, I will revise my estimate of the volume of information required to replicate a person to a whopping 15 petabytes. Although that’s 15 times my first estimate, it makes very difference to the prediction of when human replication technology will become commercial reality. Assuming Moore’s Law, a factor of 15 adds eight years. We should be able to transmit 15 PB in one second by 2057.
But it is unlikely that Moore’s Law determines the critical path for development of human replication technology. Data volume estimates give a useful measure of the size of the problem, but do not, by themselves, define the problem. They mainly help establish a floor for our expectations. Barring a wildly discontinuous breakthrough in computing technology, we can be confident we won’t see information-based human teleportation before the 2050’s. Data volume estimates do not help predict the dates of all the inventions that will be needed to solve the twin problems of recording essential information about a human organism, then constructing a human organism indistinguishable from the original: the measurement problem and the assembly problem. When a technology is well advanced, as tissue engineering is today, it is possible to get a feel for progess, to make an educated call that won’t be wrong by an order of magnitude. But when technology is at an early stage, estimating time lines becomes almost impossible.
The science of measuring brain connectivity is, I’m afraid, still at such an early stage.
The Human Connectome Project
The Human Connectome Project (HCP) aims to map the connectivity of the human brain. As described on the project website, the HCP:
…is a project to construct a map of the complete structural and functional neural connections in vivo within and across individuals. The HCP represents the first large-scale attempt to collect and share data of a scope and detail sufficient to begin the process of addressing deeply fundamental questions about human connectional anatomy and variation.
The NIH-funded research is led by two main groups. One of them, the WU-Minn consortium (Washington University in Saint Louis and the University of Minnesota), will, according to Wikipedia:
…map the connectomes in each of 1,200 healthy adults — twin pairs and their siblings from 300 families. The maps will show the anatomical and functional connections between parts of the brain for each individual, and will be related to behavioral test data. Comparing the connectomes and genetic data of genetically identical twins with fraternal twins will reveal the relative contributions of genes and environment in shaping brain circuitry and pinpoint relevant genetic variation.
The other major group, the Harvard/MGH-UCLA consortium, aims to build better instruments for brain-imaging and analysis, developing a new generation of diffusion MRI scanner expected to yield a four to eight-fold improvement over today’s technology.
The Human Connectome Project is consciously modelled on the Human Genome Project. The HGP, which achieved its main goal of recording the entire human genome by April 2003 – two years ahead of schedule – was a spectacular success. But there are differences between the connectome and the genome which make the Connectome Project more difficult.
The most important difference, I think, is that the genome is naturally in the digital domain and the connectome is not. The data elements of the genome consist of unambiguous sequences of four discrete elements, the four bases of DNA: adenine, cytosine, guanine and thymine. Because of its discrete nature, the informational content of the genome is well-defined. The genome can be expressed using exactly two bits per base. Two bits, multiplied by the number of bases (over 6 billion), gives the amount of information required to replicate the DNA of a human individual – a relatively modest 1.6 gigabytes. (A 1.6 gigabyte text file would hold 300 copies of the complete works of Shakespeare.)
The connectome is less well defined than the genome. The functional unit of the connectome is a synaptic connection between neurons. The amount of important information about a synaptic connection – what would be needed to build a functionally equivalent connection – has not been well characterized. My estimate of 176 bits per connection is only an educated guess (or if you prefer, since I have no formal training in neuroscience, a half-educated guess). The brain, unlike DNA, is in the analog domain. Some attributes essential to brain function, such as spatial proximity and signal strength, have values expressed on continuous gradients.
The fact that the brain, unlike modern electronic computers (but like many computers of the mid-20th century), is an analog device, does not mean that it cannot be replicated. But it does make it harder to know when the job is done. How close is close enough? I do not think that question has been answered yet. I will add that the answer cannot be, “Exactly the same” – that would be to set the bar far too high (perhaps to the point of meaninglessness). A living brain is not exactly the same from one second to the next; connections are constantly being made and destroyed. This fact should, eventually, allow scientists to come up with an adequate working definition of how close is close enough. The physical differences between my brain at time t and my brain at time t + 1 second can be expressed as some quantity q. If a replica made of my brain at time t differs from the original by a quantity less than, say, 0.1 q, I for one would happily accept it as “close enough.”
A second big difference between the Connectome and Genome projects is in data volumes. The difference between 1.6 GB for the genome and my estimate of 13 PB for the connectome is a factor on the order of 10 billion. A projection based on Moore’s Law – still the most reliable guide to our ability to handle data volumes – stipulates 67 years to advance from the 2003 achievement of the Genome Project to what is needed to record all connections in a human brain.
To be fair, recording all the connections in the brain is not the aim of the current Connectome Project. Goals of the 5-year HCP are limited to mapping the main conduits of brain connectivity – large bundles of axons, not individual neurons. Many future projects, as yet unplanned, will be needed to measure brain connectivity in the detail needed for human replication.
A third noteworthy difference is that, unlike the genome, the connectome is in constant flux. A 2002 study by Bonhoeffer and Yuste reports that dendritic spines frequently change shape, possibly “to enable a searching function” during the process of forming new synapses. This volatility of the connectome poses a special challenge to the goal of replicating living brains. Recording the brain’s information will take time – perhaps as little as a second, by mid-century, but still, a measurable time during which some brain connections will probably be made and others destroyed. The connectivity model resulting from that process cannot represent the brain’s state at an ‘instant’ of time; rather, it will contain information acquired at different times. Does the brain’s volatility create a risk of recording a state of the connectome that is internally inconsistent? I honestly don’t know if this is a serious concern. Perhaps if the time needed to capture the information becomes short enough, volatility will be unimportant.
Recording the Connectome – State of the Art
A good introduction to the problem of measuring brain connectivity is found in a paper titled, “MR connectomics: Principles and challenges”, by Hagmann et al. The authors compare the new field of connectomics to genomics as it was a couple of decades ago.
Noteworthy is the fact that in the early days of genomics the number and boundaries of genes was not clear either and that actually genomics technology helped to define them…. In the human cerebral cortex, neurons are arranged in an unknown number of anatomically distinct regions and areas, perhaps on the order of 100 or more. It is not clear whether cyto-architechtonically defined…or more functionally defined areas would be ideal. Nor is it clear either what the optimal scale is for efficient characterization of brain connectivity. Is it the neuronal, micro-column or the regional scale (respectively the micro-, meso- or macro-scale) that is most appropriate?
The questions that will define connectomics remain largely open. Will research be conducted on the level of neurons, axon bundles, or larger anatomical structures? Should connectomics study structural or functional units?
Much of the Hagmann paper is given to assessing the promise and limitations of diffusion imaging technology. Diffusion imaging uses magnetic resonance (MR) to measure the direction and volume of water molecules moving inside tissues. Diffusion of water indicates neuronal activity, from which the trajectories of neuron bundles and the direction of their signals can be estimated. The Diffusion Spectrum Imaging (DSI) sensor being built for the Connectome Project by the Harvard-UCLA group is based on this principle. DSI imaging cannot resolve individual neurons. Although its resolution is better than other in vivo brain scanning methods, it is not fine enough to capture the detail needed to replicate a human brain without noticeable loss of abilities, memories, and personality.
There is an older technique which does allow a connectome to be mapped to the micro-level of individual neurons and synapses – but not without destroying the brain. The MIT News of Jan 28, 2010 describes this technique used to analyze a nervous system much, much less complex than our own – that of C. elegans, a tiny worm with 302 neurons to manage its simple life. A Cambridge team needed “more than a dozen years of tedious labour” to map the complete connectome, or ‘wiring diagram,’ of those 302 neurons. Their method required slicing the nervous system into sheets, creating an image of each sheet with an electron microscope, then tracing neuronal fibres from each image to the next and identifying the connections (synapses) where neurons interact. The article describes the difficulty of applying this method even to small pieces of higher-level mammalian brains:
At the Max Planck Institute for Medical Research in Heidelberg, Germany, neuroscientists in the laboratory of Winfried Denk have assembled a team of several dozen people to manually trace connections between neurons in the retina. It’s a painstaking process — each neuron takes hours to trace, and each must be traced by as many as 10 people, in order to catch careless errors. Using this manual approach, finding the connectome of just one cubic millimeter of brain would take tens of thousands of work-years, says Viren Jain…. [emphasis added]
But progress is being made towards automating this process. A 2008 Wired article describes an “automated brain peeler and imager” called ATLUM, being developed by Harvard’s Jeff Lichtman to map the connectome of a mouse.
ATLUM uses a lathe and specialized knife to create long, thin strips of brain cells that can be imaged by an electron microscope. Software will eventually montage the images, creating an ultrahigh-resolution 3-D reconstruction of the mouse brain, allowing scientists to see features only 50 nanometers across.
“It works like an apple peeler,” Lichtman said. “Our machine takes a brain, peels off a surface layer, and puts it all on tape. These technologies will allow us to get to the finest resolution, where every single synapse is accounted for.”
Lichtman sounds a note of caution about data volumes. “A full set of images of the human brain at synapse-level resolution would contain hundreds of petabytes of information, or about the total amount of storage in Google’s data centers.” That is more than my 13-petabyte estimate – but Lichtman is talking about raw data. Those chunky 20-MB image files must be analyzed by software to create a map of the connectome, which should be considerably smaller. The analysis software cannot yet trace neural connections completely and reliably without guidance from the researchers. But repetitive image analysis and massive number-crunching are things that software is good at, and there is little doubt that a concerted effort in this direction will succeed. The Wired article says, “It could be a decade before data-crunching technology will be available to map the complexity of the human brain.” I would say five or six decades.
ATLUM does not work with a living brain. But if new imaging technology were developed which could acquire cross-sectional images of live brains at ATLUM’s 50-nanometre resolution, it’s clear that the same image-analysis software could be used to complete the job of building the connectome. Since new brain imaging techniques seem to come along every two or three years, that is not too much to hope for.
Building a Brain
A solution to the assembly problem is further off and harder to see than a solution to the measurement problem. I don’t know of any research that has even begun in this area. It seems clear that the bioprinting techniques which hold so much promise for the rest of the body could not be used to reproduce a human brain with all its connections. A single neuron may be up to a metre long; its axon may have multiple branches, and its dendrites may terminate at thousands of synaptic connections with other neurons. It is hard to imagine a bioprinter reproducing those connections by spraying neurons from a print head, however precise. The three-dimensional neural web could not be physically reproduced by such a layered technique. It seems more likely that the neurons themselves will have to be ‘printed’, a slice at a time. That calls for an assembly process that works at the molecular, not the cellular level. It will require keeping the neurons ‘alive’ – somehow potentially viable – during their construction, so that they will be truly alive when completed. In 2010, it is hard to know when that might become possible.
But one thing we know about the future is that – barring massive catastrophes –unforeseen and unforeseeable progress will be made. Fifty years ago, when I was twelve, my wife’s iPad, with its internet connection, would have seemed magical – not metaphorically, but literally magical. Computers in 1960 had vacuum tubes and punched-card readers; they occupied large clean-rooms, and output text to line printers. The technical advances of the past fifty years could not have been guessed at, let alone understood, by someone with a 1960’s engineering background trying to comprehend how an iPad responds to questions typed into a Google search box. If I’d witnessed a working, internet-connected iPad in 1960, I couldn’t have done much better than, “How about that – crystal balls are flat!”
References
Azevedo, FA et al (2009), “Equal numbers of neuronal and nonneuronal cells make the human brain an isometrically scaled-up primate brain”. J Comp Neurol. 2009 Apr 10;513(5):532-41.
Bonhoeffer, T and Yuste, R (2002), “Spine Motility”. Neuron, Volume 35, Issue 6, 1019-1027, 12 September 2002
Drachman, David (2005), “Do We Have Brain to Spare?”. Neurology (2005;64:2004-5).
Hagmann P, et al (2010), “MR connectomics: Principles and challenges”. J Neurosci Methods (2010), doi:10.1016/j.jneumeth.2010.01.014

Given everything you have said, it still remains an interesting topic the one of personhood replication. Compared to the relevance of the mere phenomenological quality of personhood, however, it remains as a rather secondary concern. What seems to really matter, as I can follow from your earlier discussions, is the weight you can put into a completely apersonal phenomena.
Thanks for bringing up the phenomenological quality of personhood. This is a subject I’d like to explore. I’ve been reading Thomas Metzinger’s Being No One – quite a difficult book, but rewarding if you stick with it. I hope to have something to say about it before too long.